import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
# Fine source field
lats_f = np.arange(25, 50, 0.5)
lons_f = np.arange(-120, -70, 0.5)
LON, LAT = np.meshgrid(lons_f, lats_f)
T = 300 - 0.5*(LAT - 25) + 3*np.sin(np.radians(LON+100))
T += np.random.default_rng(42).normal(0, 0.5, T.shape)
ds = xr.Dataset({'t2m': (['lat','lon'], T)}, coords={'lat': lats_f, 'lon': lons_f})
lats_c = np.arange(26, 50, 2.0)
lons_c = np.arange(-119, -70, 2.0)
ds_nn = ds.sel(lat=lats_c, lon=lons_c, method='nearest')
ds_bil = ds.interp(lat=lats_c, lon=lons_c)
# Cross-section along a fixed latitude
lat_xs = float(ds['t2m'].sel(lat=38.0, method='nearest').lat)
fine_xs = ds['t2m'].sel(lat=lat_xs, method='nearest')
nn_xs = ds_nn['t2m'].sel(lat=lat_xs, method='nearest')
bil_xs = ds_bil['t2m'].sel(lat=lat_xs, method='nearest')
from matplotlib.gridspec import GridSpec
fig = plt.figure(figsize=(13, 7))
gs = GridSpec(2, 3, figure=fig, hspace=0.45, wspace=0.05,
top=0.93, bottom=0.08, left=0.06, right=0.94)
ax_maps = [fig.add_subplot(gs[0, i]) for i in range(3)]
ax_xs = fig.add_subplot(gs[1, :])
vmin, vmax = float(ds['t2m'].min()), float(ds['t2m'].max())
for ax, data, title in zip(ax_maps,
[ds['t2m'], ds_nn['t2m'], ds_bil['t2m']],
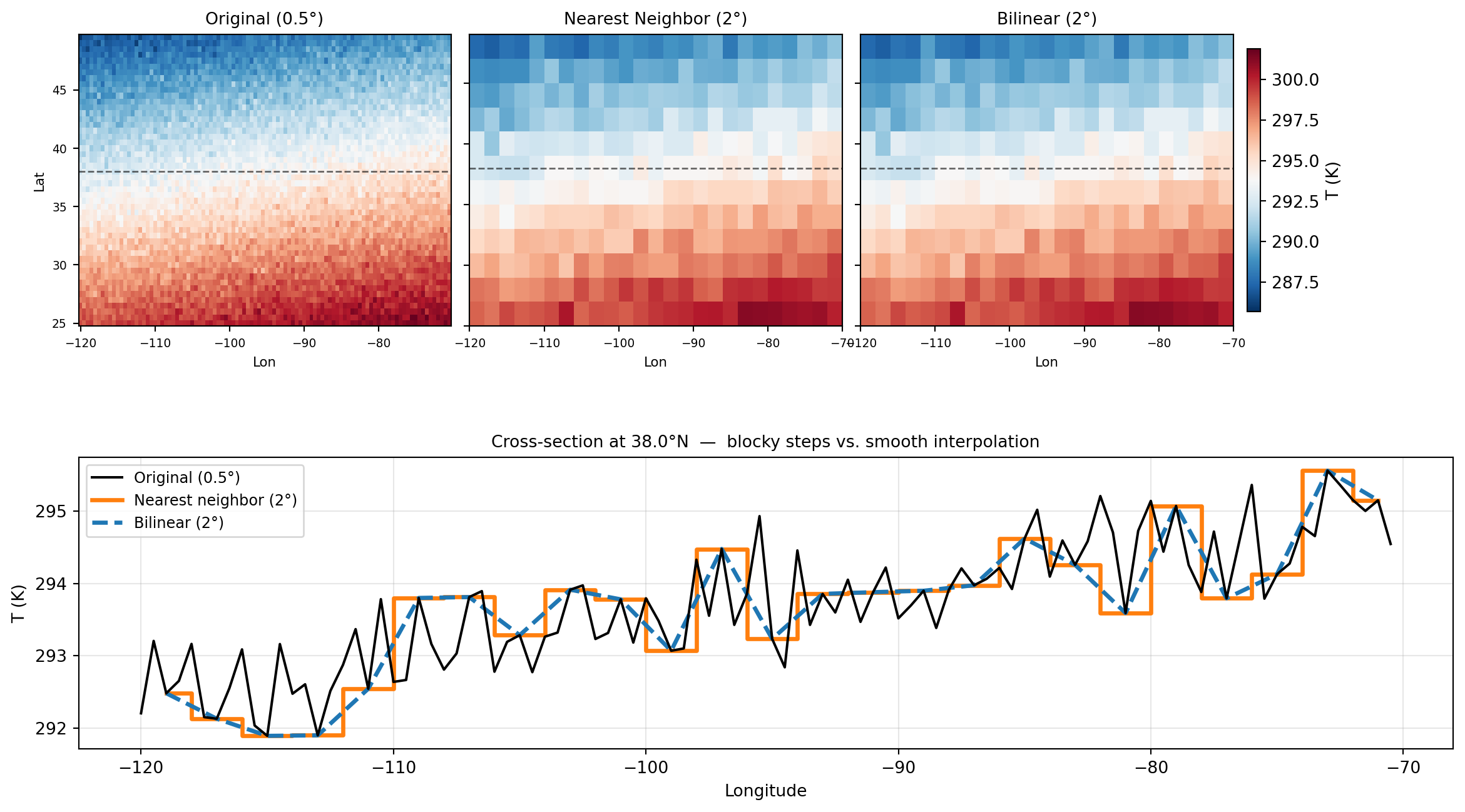
['Original (0.5°)', 'Nearest Neighbor (2°)', 'Bilinear (2°)']):
im = ax.pcolormesh(data.lon, data.lat, data.values,
vmin=vmin, vmax=vmax, cmap='RdBu_r')
ax.set_title(title, fontsize=10)
ax.set_xlabel('Lon', fontsize=8)
ax.tick_params(labelsize=7)
ax.axhline(lat_xs, color='k', lw=1, ls='--', alpha=0.6)
ax_maps[0].set_ylabel('Lat', fontsize=8)
for ax in ax_maps[1:]:
ax.set_yticklabels([])
fig.colorbar(im, ax=ax_maps, shrink=0.9, pad=0.01, label='T (K)')
# Cross-section: the key comparison
ax_xs.plot(fine_xs.lon, fine_xs.values, 'k-', lw=1.5, label='Original (0.5°)', zorder=3)
ax_xs.step(nn_xs.lon, nn_xs.values, where='mid',
color='tab:orange', lw=2.5, label='Nearest neighbor (2°)')
ax_xs.plot(bil_xs.lon, bil_xs.values, '--', lw=2.5,
color='tab:blue', label='Bilinear (2°)', zorder=2)
ax_xs.set_xlabel('Longitude')
ax_xs.set_ylabel('T (K)')
ax_xs.set_title(f'Cross-section at {lat_xs:.1f}°N — blocky steps vs. smooth interpolation',
fontsize=10)
ax_xs.legend(fontsize=9)
ax_xs.grid(alpha=0.3)
plt.savefig('/tmp/regrid_comparison.png', dpi=100, bbox_inches='tight')
plt.show()