Mon 15.2
Tue 18.7
Wed 22.1
Thu 19.8
dtype: float64ATOC 4815/5815
Tabular Data & Pandas Foundations - Week 7
CU Boulder ATOC
Spring 2026
Reminders
Due this evening at 12pm:
- Lab 4
Office Hours:
Will: Tu 11:15-12:15p; Th 9-9:50a
Aerospace Cafe
Aiden: M / W 330-430p
DUAN D319
ATOC 4815/5815 Playlist

Tabular Data & Pandas
Today’s Objectives
- Understanding pandas: from arrays to tables
- Reading and parsing CSV files with dates
- Time series indexing and resampling
- Rolling windows and aggregations
- Creating publication-quality time series plots
- Recognizing and fixing common pandas errors
- Building analysis workflows you’ll use in research
The Real Problem
Your Research Scenario
Imagine: You’re analyzing Boulder’s urban heat island effect
Your data:
- 10 ASOS weather stations around Boulder
- 1 year of hourly measurements (8,760 hours × 10 stations = 87,600 rows!)
- Multiple variables: temperature, humidity, wind speed, pressure, precipitation
Each station CSV looks like:
Date and Time,Station,Temp_C,RH_pct,Wind_kt,Pressure_hPa,Precip_mm
2024-01-01 00:00,KBDU,2.1,65,8,1013.2,0.0
2024-01-01 01:00,KBDU,1.8,68,7,1013.5,0.0
2024-01-01 02:00,KBDU,1.2,71,6,1013.8,0.2
...Questions you need to answer:
- What’s the average daily temperature at each station?
- Which station is warmest? When?
- How does precipitation accumulate over the month?
- Are there heat waves (3+ consecutive days > 30°C)?
Why NumPy Arrays Fall Short
Try solving this with NumPy arrays…
Problem 1: Multiple data types
Problem 2: No column names
What Pandas Gives Us
Pandas solves all these problems:
1. Mixed data types in columns:
2. Named columns:
3. Time-aware operations:
Bottom line: For tabular data with time series, Pandas is the right tool. NumPy is for uniform numeric arrays and math.
Why Pandas?
Pandas Scientific Story
Created by Wes McKinney (late 2000s) to handle panel data for quantitative finance
- Goal: bring R/Excel/SQL-style table tools into Python
Built on top of NumPy, adding:
- Labeled rows/columns (
DataFrame,Series) - Easy handling of missing values
- Powerful time-series tools (date indexes, resampling, rolling windows)
Became the standard tabular data library in scientific Python:
- Most data tutorials start with
import pandas as pd - Common front-end for reading/writing CSV, Excel, SQL, NetCDF/Parquet, etc.
- Feeds directly into NumPy, Matplotlib, and higher-level tools (xarray, geopandas, statsmodels)
For this course, think of pandas as:
“Excel + SQL + NumPy, but in code” — a single place to clean data, compute statistics, and drive time-series visualizations.
Mental Model: NumPy vs Pandas
Think of it this way:
NumPy: "Calculator for arrays of numbers"
✅ Fast math, vectorized operations
❌ No column names, no mixed types, weak time handling
Pandas: "Spreadsheet + database in Python"
✅ Named columns, mixed types, time series tools
✅ Built on NumPy (uses arrays internally)
❌ Slightly slower (but worth it for convenience)Use NumPy when:
- Doing heavy numerical computation (matrix ops, FFT, stats)
- All data is numeric and uniform
Use Pandas when:
- Working with tables (CSV, Excel, SQL)
- Have mixed data types (strings, dates, numbers)
- Need time-based operations (resampling, rolling windows)
- Want readable code with named columns
Check Your Understanding
Which tool should you use for each task?
1. Computing the FFT of 10,000 temperature measurements
Answer: NumPy (uniform numeric array, pure math operation)
2. Loading a CSV with station names, timestamps, temps, and wind speeds
Answer: Pandas (mixed types, labeled columns, time data)
3. Calculating daily mean temperature from hourly data
Answer: Pandas (time-based resampling with .resample('1D').mean())
4. Multiplying two 1000×1000 matrices
Answer: NumPy (pure numeric computation with np.dot() or @)
Pandas Fundamentals
Series: One Column
A Series is a 1-D labeled array (like one column of a spreadsheet)
Key features:
.index→ row labels (Mon, Tue, Wed, Thu).values→ underlying NumPy array- Access by label:
temps['Mon']→ 15.2 - Access by position:
temps[0]→ 15.2
DataFrame: Multiple Series
A DataFrame is a 2-D table (like a whole spreadsheet)
temp_c pressure_hpa
Mon 15.2 1010
Tue 18.7 1012
Wed 22.1 1008
Thu 19.8 1011Key features:
.columns→ column names (temp_c, pressure_hpa).index→ row labels (Mon, Tue, Wed, Thu)- Each column is a Series
- Access column:
data['temp_c']→ Series - Access row:
data.loc['Mon']→ Series
Common Error: Accessing Columns
Predict the output:
0 15.2
1 18.7
2 22.1
Name: temp_c, dtype: float64** It works!** But there’s a catch…
Common Error: KeyError
Predict the output:
KeyError: 'temperature'Explanation: Column ‘temperature’ doesn’t exist (it’s ‘temp_c’)
The Fix:
Common causes:
- Typo in column name
- Wrong capitalization (‘Temp_C’ vs ‘temp_c’)
- Column was renamed or not read from CSV
Try It Yourself 💻
With your neighbor (3 min): Create a DataFrame with Boulder weather
weather = pd.DataFrame({
'date': ['2024-01-01', '2024-01-02', '2024-01-03'],
'temp_c': [2.1, 3.5, 1.2],
'precip_mm': [0.0, 2.5, 0.5]
})
# Tasks:
# 1. Print the DataFrame
# 2. Extract just the temp_c column
# 3. What's the maximum precipitation?
# 4. Try accessing a column that doesn't exist—what error do you get?Reading Data
Reading CSVs: The Hard Way
What happens if you just read the CSV naively?
# Sample CSV as a string (simulating a file)
import io
csv_data = """Date and Time,Station,Temp_C
2024-01-01 00:00,KBDU,2.1
2024-01-01 01:00,KBDU,1.8
2024-01-01 02:00,KBDU,1.2"""
df_wrong = pd.read_csv(io.StringIO(csv_data))
print(df_wrong)
print(f"\nData type of 'Date and Time': {df_wrong['Date and Time'].dtype}") Date and Time Station Temp_C
0 2024-01-01 00:00 KBDU 2.1
1 2024-01-01 01:00 KBDU 1.8
2 2024-01-01 02:00 KBDU 1.2
Data type of 'Date and Time': objectProblem: ‘Date and Time’ is stored as a string (object), not a timestamp!
Why it matters:
- Can’t do time-based operations (resampling, rolling windows)
- Can’t filter by date easily
- Can’t extract month, day, hour
Reading CSVs: The Easy Way
Use parse_dates to convert string → datetime:
Date and Time Station Temp_C
0 2024-01-01 00:00:00 KBDU 2.1
1 2024-01-01 01:00:00 KBDU 1.8
2 2024-01-01 02:00:00 KBDU 1.2
Data type of 'Date and Time': datetime64[ns]Now it’s a datetime64 type!
What you can do now:
Common Error: Forgetting parse_dates
Predict the output:
TypeError: Only valid with DatetimeIndex, TimedeltaIndex or PeriodIndex,
but got an instance of 'RangeIndex'Explanation: Can’t resample without a time index!
The Fix:
# Method 1: Parse on read
df = pd.read_csv('weather.csv', parse_dates=['Date and Time'])
df = df.set_index('Date and Time')
daily = df.resample('1D').mean() # ✅ Works!
# Method 2: Convert after reading
df = pd.read_csv('weather.csv')
df['Date and Time'] = pd.to_datetime(df['Date and Time'])
df = df.set_index('Date and Time')Setting a Time Index
For time series analysis, make the timestamp the index
Why?
- Enables
.resample(),.rolling(), time-based slicing - Aligns operations by time automatically
- Makes plots use time on x-axis by default
temp_c pressure_hpa
2024-01-01 00:00:00 15.2 1010
2024-01-01 01:00:00 16.1 1011
2024-01-01 02:00:00 17.3 1009
2024-01-01 03:00:00 18.2 1008
2024-01-01 04:00:00 17.5 1010
Index type: <class 'pandas.core.indexes.datetimes.DatetimeIndex'>Setting Index: Two Methods
Method 1: Set index after reading
Method 2: Set index during read (more efficient)
Method 2 is better:
- One step instead of two
- Slightly faster (one less copy)
- Less code to maintain
Resampling & Aggregation
What is Resampling?
Resampling: Change the frequency of your time series
Visual example:
Hourly data (24 points per day):
├─ 00:00 → 15.2°C
├─ 01:00 → 16.1°C
├─ 02:00 → 17.3°C
├─ 03:00 → 18.2°C
...
Resample to daily (1 point per day):
└─ 2024-01-01 → 16.7°C (mean of all 24 hours)Common patterns:
- Downsampling: High → Low frequency (hourly → daily)
- Aggregation: How to combine values (mean, sum, max, min, etc.)
Resampling Syntax
# Create 15-minute data
dates = pd.date_range('2024-01-01', periods=96, freq='15min')
df = pd.DataFrame({
'temp_c': 15 + 5 * np.sin(np.arange(96) * 2 * np.pi / 96) + np.random.randn(96) * 0.5,
'precip_mm': np.random.exponential(0.1, 96)
}, index=dates)
print("Original (15-min):")
print(df.head())
# Resample to hourly
hourly = df.resample('1h').mean()
print("\nResampled (hourly):")
print(hourly.head())Original (15-min):
temp_c precip_mm
2024-01-01 00:00:00 15.679592 0.021174
2024-01-01 00:15:00 15.882105 0.005647
2024-01-01 00:30:00 15.485879 0.104231
2024-01-01 00:45:00 16.991130 0.028296
2024-01-01 01:00:00 17.012686 0.000142
Resampled (hourly):
temp_c precip_mm
2024-01-01 00:00:00 16.009676 0.039837
2024-01-01 01:00:00 16.642454 0.061717
2024-01-01 02:00:00 17.659040 0.061184
2024-01-01 03:00:00 18.528682 0.035272
2024-01-01 04:00:00 19.519600 0.159394Aggregation Rules: When to Use What?
Different variables need different aggregation methods:
| Variable | Aggregation | Why? |
|---|---|---|
| Temperature | mean() |
Average temp over period makes sense |
| Precipitation | sum() |
Want total accumulated precip |
| Wind speed | mean() or max() |
Mean for typical, max for gusts |
| Pressure | mean() |
Average pressure over period |
| Station ID | first() |
Metadata—just keep one |
Example:
temp_c precip_mm
2024-01-01 00:00:00 16.009676 0.159347
2024-01-01 01:00:00 16.642454 0.246869
2024-01-01 02:00:00 17.659040 0.244737Common Error: Wrong Aggregation
Predict the problem:
The Problem: Using mean() for precipitation!
- You don’t want the “average hourly precip”
- You want “total daily precip”
The Fix:
Example:
Hourly: [0.5, 0.2, 0.0, 0.8] mm
Daily (wrong): mean = 0.375 mm ← What does this even mean?
Daily (right): sum = 1.5 mm ← Total precip for the dayTry It Yourself 💻
With your neighbor (5 min): Practice resampling
# Create hourly temperature data
dates = pd.date_range('2024-01-01', periods=168, freq='h') # 1 week
temps = pd.Series(
15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168),
index=dates
)
# Tasks:
# 1. Resample to daily mean temperature
# 2. Find the warmest day
# 3. Resample to 6-hour max temperature
# 4. What happens if you resample but forget to call .mean() or .sum()?Answers:
# 1. Daily mean
daily = temps.resample('1D').mean()
# 2. Warmest day
warmest = daily.idxmax() # Returns the date
print(f"Warmest day: {warmest} at {daily.max():.1f}°C")
# 3. 6-hour max
six_hour_max = temps.resample('6h').max()
# 4. Forget aggregation
resampled = temps.resample('1D') # Just returns a Resampler object, not data!
print(resampled) # DatetimeIndexResampler [freq=<Day>, ...]Multiple Aggregations
You can compute multiple statistics at once:
# Create sample data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
df = pd.DataFrame({
'temp_c': 15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
'precip_mm': np.random.exponential(0.3, 168)
}, index=dates)
# Daily aggregation with different rules
daily = df.resample('1D').agg({
'temp_c': ['mean', 'min', 'max'],
'precip_mm': 'sum'
})
print(daily) temp_c precip_mm
mean min max sum
2024-01-01 14.445156 5.937024 24.848040 7.662438
2024-01-02 14.745927 2.868327 26.082912 6.986567
2024-01-03 15.326128 5.066317 23.055476 5.987383
2024-01-04 15.597366 5.220643 24.748672 5.374198
2024-01-05 14.871517 4.761499 24.987079 5.766746
2024-01-06 15.425050 5.373944 24.486201 7.337340
2024-01-07 15.008559 3.895228 24.674200 6.478123Accessing multi-level columns:
First day mean temp: 14.4°C
First day total precip: 7.66 mmResampling Frequency Codes
Common frequency strings:
| Code | Meaning | Example |
|---|---|---|
'1h' |
Hourly | Every hour |
'3h' |
Every 3 hours | 00:00, 03:00, 06:00, … |
'1D' |
Daily | Once per day |
'1W' |
Weekly | Once per week |
'1MS' |
Monthly (start) | First day of each month |
'1ME' |
Monthly (end) | Last day of each month |
'1QS' |
Quarterly (start) | Jan 1, Apr 1, Jul 1, Oct 1 |
'1YS' |
Yearly (start) | Jan 1 each year |
Rolling Windows
What is a Rolling Window?
Rolling window: Compute statistics over a moving time window
Visual example:
Data: [10, 12, 15, 18, 20, 22, 21, 19, 16, 14]
↓ ↓ ↓
Window: [10, 12, 15] → mean = 12.3
↓ ↓ ↓
Window: [12, 15, 18] → mean = 15.0
↓ ↓ ↓
Window: [15, 18, 20] → mean = 17.7
...Result: A smoothed version of the original data
Rolling Window Syntax
# Create hourly temperature data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
temps = pd.Series(
15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
index=dates
)
# Rolling means with different windows
temps_3h = temps.rolling('3h').mean()
temps_12h = temps.rolling('12h').mean()
temps_24h = temps.rolling('24h').mean()
print("Original vs Rolling means:")
df_compare = pd.DataFrame({
'original': temps,
'rolling_3h': temps_3h,
'rolling_12h': temps_12h,
'rolling_24h': temps_24h
})
print(df_compare.head(26))Original vs Rolling means:
original rolling_3h rolling_12h rolling_24h
2024-01-01 00:00:00 14.262789 14.262789 14.262789 14.262789
2024-01-01 01:00:00 18.787965 16.525377 16.525377 16.525377
2024-01-01 02:00:00 20.731017 17.927257 17.927257 17.927257
2024-01-01 03:00:00 17.585874 19.034952 17.841911 17.841911
2024-01-01 04:00:00 25.658387 21.325093 19.405206 19.405206
2024-01-01 05:00:00 24.972099 22.738787 20.333022 20.333022
2024-01-01 06:00:00 22.554884 24.395123 20.650431 20.650431
2024-01-01 07:00:00 26.028218 24.518400 21.322654 21.322654
2024-01-01 08:00:00 21.684229 23.422444 21.362829 21.362829
2024-01-01 09:00:00 20.948094 22.886847 21.321356 21.321356
2024-01-01 10:00:00 18.447893 20.360072 21.060132 21.060132
2024-01-01 11:00:00 17.489995 18.961994 20.762620 20.762620
2024-01-01 12:00:00 17.655007 17.864298 21.045305 20.523573
2024-01-01 13:00:00 7.612165 14.252389 20.113988 19.601330
2024-01-01 14:00:00 9.716695 11.661289 19.196128 18.942354
2024-01-01 15:00:00 6.280261 7.869707 18.253994 18.150973
2024-01-01 16:00:00 8.403746 8.133567 16.816107 17.577607
2024-01-01 17:00:00 4.695333 6.459780 15.126377 16.861925
2024-01-01 18:00:00 5.147387 6.082155 13.675752 16.245370
2024-01-01 19:00:00 6.528044 5.456921 12.050737 15.759504
2024-01-01 20:00:00 9.479993 7.051808 11.033718 15.460480
2024-01-01 21:00:00 8.428825 8.145621 9.990445 15.140859
2024-01-01 22:00:00 8.386323 8.765047 9.151981 14.847184
2024-01-01 23:00:00 11.310491 9.375213 8.637022 14.699821
2024-01-02 00:00:00 18.706638 12.801151 8.724658 14.884982
2024-01-02 01:00:00 14.822106 14.946412 9.325487 14.719738Rolling Window Visualization
import matplotlib.pyplot as plt
# Plot raw data and rolling means
plt.figure(figsize=(9, 4))
plt.plot(temps.index, temps, alpha=0.3, label='Raw (hourly)', linewidth=1)
plt.plot(temps.index, temps_3h, label='3-hour rolling mean', linewidth=1.5)
plt.plot(temps.index, temps_12h, label='12-hour rolling mean', linewidth=1.5)
plt.plot(temps.index, temps_24h, label='24-hour rolling mean', linewidth=2)
plt.xlabel('Date')
plt.ylabel('Temperature (°C)')
plt.title('Temperature with Rolling Means')
plt.legend()
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()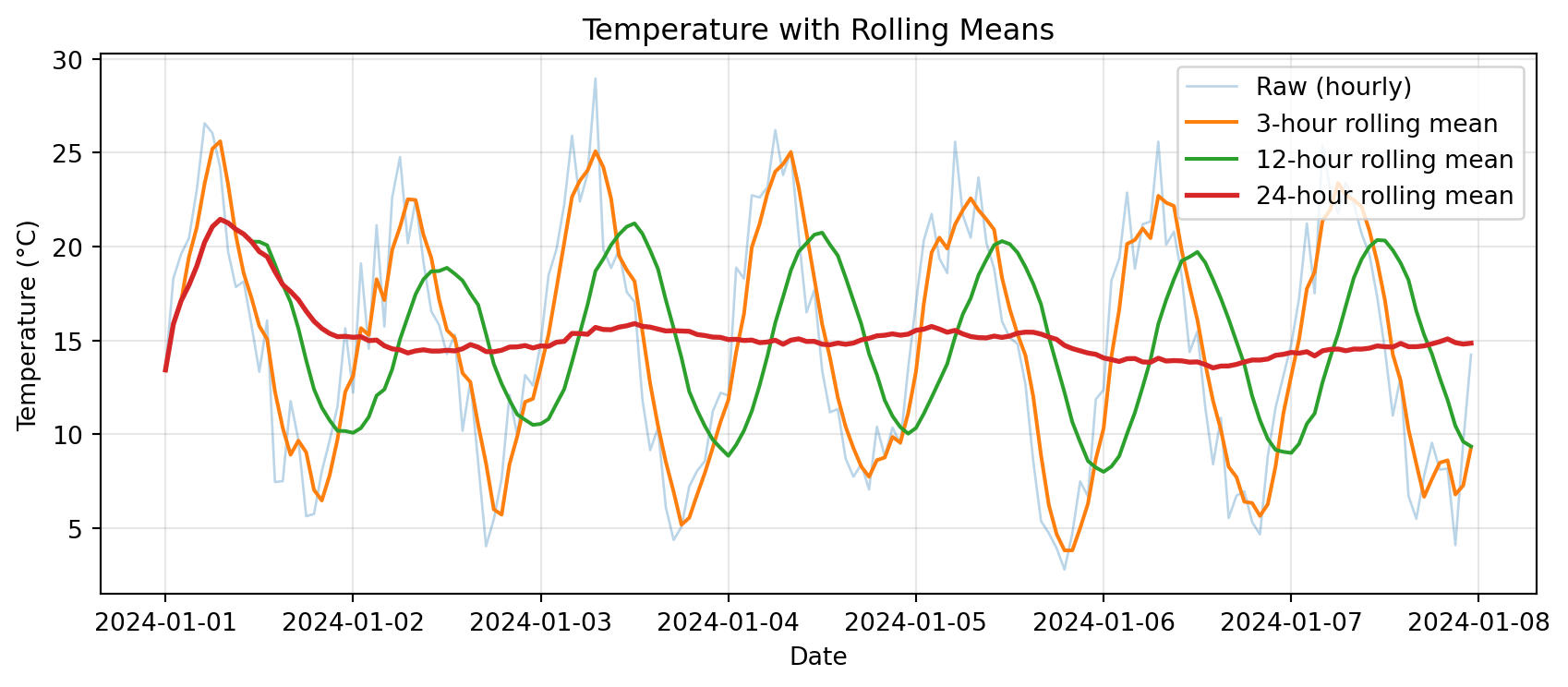
Notice: Longer windows → smoother curves, but more lag
Rolling vs Resampling: What’s the Difference?
Resampling: Change frequency (hourly → daily)
- Reduces number of points
- Each point represents a whole period
Rolling: Smooth data over a moving window
- Same number of points (except edge NaNs)
- Each point is an average of nearby points
Example:
# Original: 168 hourly points
print(f"Original: {len(temps)} points")
# Resampling to daily: reduces to 7 points
daily = temps.resample('1D').mean()
print(f"Resampled daily: {len(daily)} points")
# Rolling 24-hour: still 168 points (but first 23 are NaN)
rolling_24h = temps.rolling('24h').mean()
print(f"Rolling 24h: {len(rolling_24h)} points (includes NaN at start)")Original: 168 points
Resampled daily: 7 points
Rolling 24h: 168 points (includes NaN at start)Rolling Statistics Beyond Mean
Rolling windows aren’t just for means:
# Create temperature data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
temps = pd.Series(
15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
index=dates
)
# Different rolling statistics
rolling_mean = temps.rolling('24h').mean()
rolling_std = temps.rolling('24h').std()
rolling_min = temps.rolling('24h').min()
rolling_max = temps.rolling('24h').max()
print("Rolling 24-hour statistics:")
print(pd.DataFrame({
'mean': rolling_mean,
'std': rolling_std,
'min': rolling_min,
'max': rolling_max
}).describe())Rolling 24-hour statistics:
mean std min max
count 168.000000 167.000000 168.000000 168.000000
mean 15.338181 5.738652 6.221530 24.141977
std 1.247872 0.870020 2.820697 1.179792
min 14.474858 0.262933 4.026665 15.497366
25% 14.780460 5.609293 4.597806 23.628852
50% 14.971259 5.976296 5.339147 24.109885
75% 15.255654 6.223449 6.933531 24.759307
max 20.703824 6.600963 15.497366 25.246161Use Case: Temperature Variability
Rolling standard deviation shows how variable conditions are:
# Create data with changing variability
dates = pd.date_range('2024-01-01', periods=168, freq='h')
# Add more noise in second half
noise = np.concatenate([
np.random.randn(84) * 1, # Low variability
np.random.randn(84) * 4 # High variability
])
temps = pd.Series(
15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + noise,
index=dates
)
rolling_mean = temps.rolling('12h').mean()
rolling_std = temps.rolling('12h').std()
# Plot
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(9, 5), sharex=True)
ax1.plot(temps.index, temps, alpha=0.4, label='Hourly temp')
ax1.plot(temps.index, rolling_mean, linewidth=2, label='12-h mean')
ax1.set_ylabel('Temperature (°C)')
ax1.set_title('Temperature and Variability')
ax1.legend()
ax1.grid(True, alpha=0.3)
ax2.plot(temps.index, rolling_std, color='red', linewidth=2)
ax2.set_xlabel('Date')
ax2.set_ylabel('Rolling Std Dev (°C)')
ax2.set_title('12-hour Rolling Variability')
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()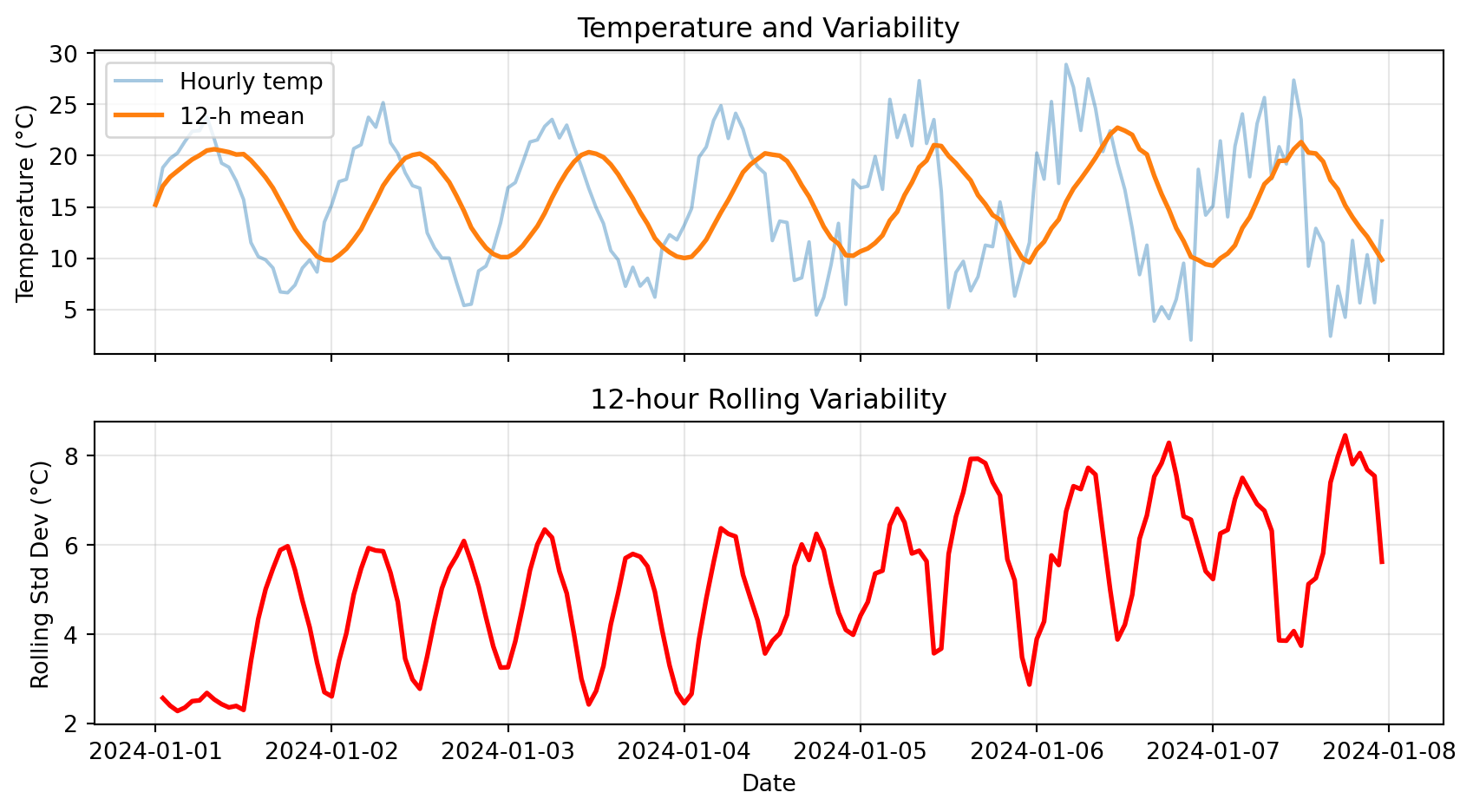
Notice: High rolling std in second half → more variable conditions
Try It Yourself 💻
With your neighbor (5 min): Explore rolling windows
# Create 1 week of hourly wind speed data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
wind_kt = pd.Series(
10 + 5 * np.abs(np.sin(np.arange(168) * 2 * np.pi / 24)) + np.random.randn(168) * 2,
index=dates
)
# Tasks:
# 1. Compute 6-hour rolling mean wind speed
# 2. Find the time period with highest 6-hour average wind
# 3. Compute 12-hour rolling max (for peak gusts)
# 4. What's the difference between .rolling(6) and .rolling('6h')?Answers:
# 1. 6-hour rolling mean
rolling_mean_6h = wind_kt.rolling('6h').mean()
# 2. Highest 6-hour average
max_time = rolling_mean_6h.idxmax()
max_wind = rolling_mean_6h.max()
print(f"Highest 6-h avg wind: {max_wind:.1f} kt at {max_time}")
# 3. 12-hour rolling max
rolling_max_12h = wind_kt.rolling('12h').max()
# 4. Difference:
# .rolling(6) → 6 data points (may not be 6 hours if data is irregular)
# .rolling('6h') → 6 hours of data (time-aware, handles gaps correctly)Anomalies & Cumulative Sums
Computing Anomalies
Anomaly: Deviation from a baseline (climatology, daily mean, etc.)
Why anomalies matter:
- Identify unusual events
- Remove seasonal cycle
- Compare different stations or years
Example: Hourly temperature anomalies from daily mean
# Create hourly data for a week
dates = pd.date_range('2024-01-01', periods=168, freq='h')
temps = pd.Series(
15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
index=dates
)
df = pd.DataFrame({'temp_c': temps})
# Method: use groupby to compute daily mean, then subtract
df['date'] = df.index.date
df['daily_mean'] = df.groupby('date')['temp_c'].transform('mean')
df['anomaly'] = df['temp_c'] - df['daily_mean']
print(df[['temp_c', 'daily_mean', 'anomaly']].head(10)) temp_c daily_mean anomaly
2024-01-01 00:00:00 16.002414 14.945173 1.057242
2024-01-01 01:00:00 19.738942 14.945173 4.793770
2024-01-01 02:00:00 16.149205 14.945173 1.204032
2024-01-01 03:00:00 17.444251 14.945173 2.499078
2024-01-01 04:00:00 19.412783 14.945173 4.467610
2024-01-01 05:00:00 22.067586 14.945173 7.122413
2024-01-01 06:00:00 23.423461 14.945173 8.478289
2024-01-01 07:00:00 24.596492 14.945173 9.651320
2024-01-01 08:00:00 24.069670 14.945173 9.124498
2024-01-01 09:00:00 19.590393 14.945173 4.645220Visualizing Anomalies
# Plot raw data and anomalies
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(9, 5), sharex=True)
# Top: Raw temperature
ax1.plot(df.index, df['temp_c'], label='Hourly temp', alpha=0.7)
ax1.plot(df.index, df['daily_mean'], label='Daily mean', linewidth=2, color='red')
ax1.set_ylabel('Temperature (°C)')
ax1.set_title('Raw Temperature and Daily Mean')
ax1.legend()
ax1.grid(True, alpha=0.3)
# Bottom: Anomalies
ax2.plot(df.index, df['anomaly'], color='purple', alpha=0.7)
ax2.axhline(y=0, color='black', linestyle='--', linewidth=1)
ax2.set_xlabel('Date')
ax2.set_ylabel('Anomaly (°C)')
ax2.set_title('Hourly Anomalies from Daily Mean')
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()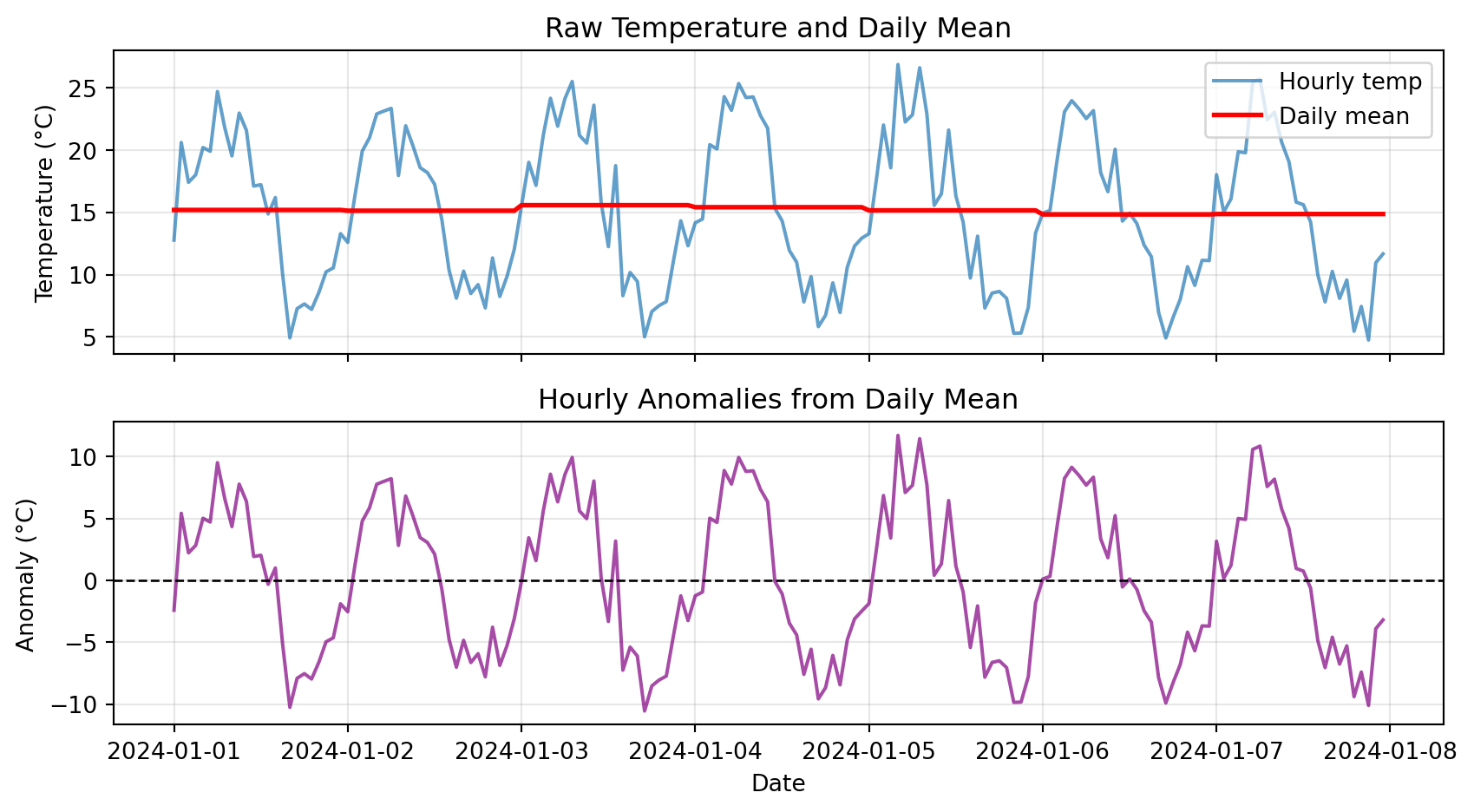
Notice: Anomalies oscillate around zero, showing deviations from typical pattern
Cumulative Sums
Cumulative sum: Running total over time
Use cases:
- Precipitation: Total accumulated rainfall
- Energy: Cumulative power consumption
- Degree-days: Accumulated heat or cold
# Create precipitation data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
precip = pd.Series(np.random.exponential(0.3, 168), index=dates)
# Compute cumulative sum
cumulative = precip.cumsum()
print("Hourly and cumulative precipitation:")
print(pd.DataFrame({
'hourly_mm': precip,
'cumulative_mm': cumulative
}).head(10))Hourly and cumulative precipitation:
hourly_mm cumulative_mm
2024-01-01 00:00:00 0.116316 0.116316
2024-01-01 01:00:00 0.032834 0.149150
2024-01-01 02:00:00 0.152598 0.301747
2024-01-01 03:00:00 0.819951 1.121699
2024-01-01 04:00:00 0.164993 1.286692
2024-01-01 05:00:00 0.121761 1.408453
2024-01-01 06:00:00 0.020566 1.429018
2024-01-01 07:00:00 0.067645 1.496663
2024-01-01 08:00:00 0.406293 1.902956
2024-01-01 09:00:00 0.103481 2.006438Cumulative Precipitation Visualization
# Plot hourly and cumulative
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(9, 5), sharex=True)
# Top: Hourly bars
ax1.bar(precip.index, precip, width=0.04, alpha=0.6, color='steelblue')
ax1.set_ylabel('Hourly Precip (mm)')
ax1.set_title('Hourly Precipitation')
ax1.grid(True, alpha=0.3)
# Bottom: Cumulative line
ax2.plot(cumulative.index, cumulative, linewidth=2, color='darkblue')
ax2.set_xlabel('Date')
ax2.set_ylabel('Cumulative Precip (mm)')
ax2.set_title('Cumulative Precipitation')
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print(f"\nTotal precipitation over week: {cumulative.iloc[-1]:.2f} mm")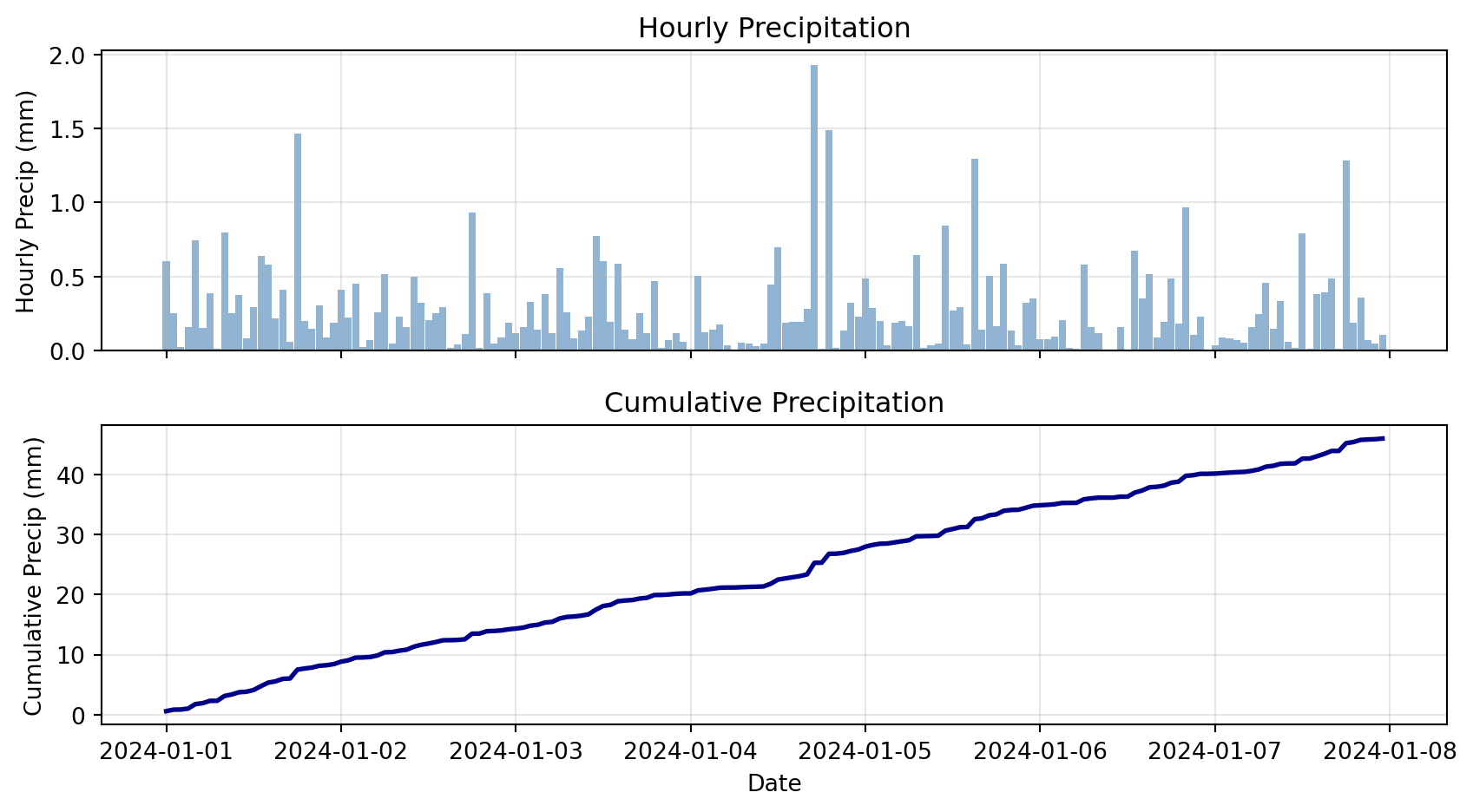
Total precipitation over week: 50.02 mmCheck Your Understanding
Match each technique to its use case:
Techniques:
- Rolling mean
- Resampling
- Anomaly
- Cumulative sum
Use cases:
A. “How much total rainfall since Jan 1?”
B. “What’s the smoothed temperature trend?”
C. “Convert hourly data to daily averages”
D. “How much warmer than normal was today?”
Answers:
1-B (Rolling mean → smoothing)
2-C (Resampling → change frequency)
3-D (Anomaly → deviation from baseline)
4-A (Cumulative sum → total accumulated)
Plotting Time Series
Pandas Native Plotting
Pandas DataFrames have built-in .plot() method:
# Create sample data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
df_plot = pd.DataFrame({
'temp_c': 15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
}, index=dates)
df_plot['rolling_24h'] = df_plot['temp_c'].rolling('24h').mean()
# Simple plot
df_plot.plot(
figsize=(9, 4),
title='Temperature Time Series',
ylabel='Temperature (°C)',
grid=True,
alpha=0.7
)
plt.tight_layout()
plt.show()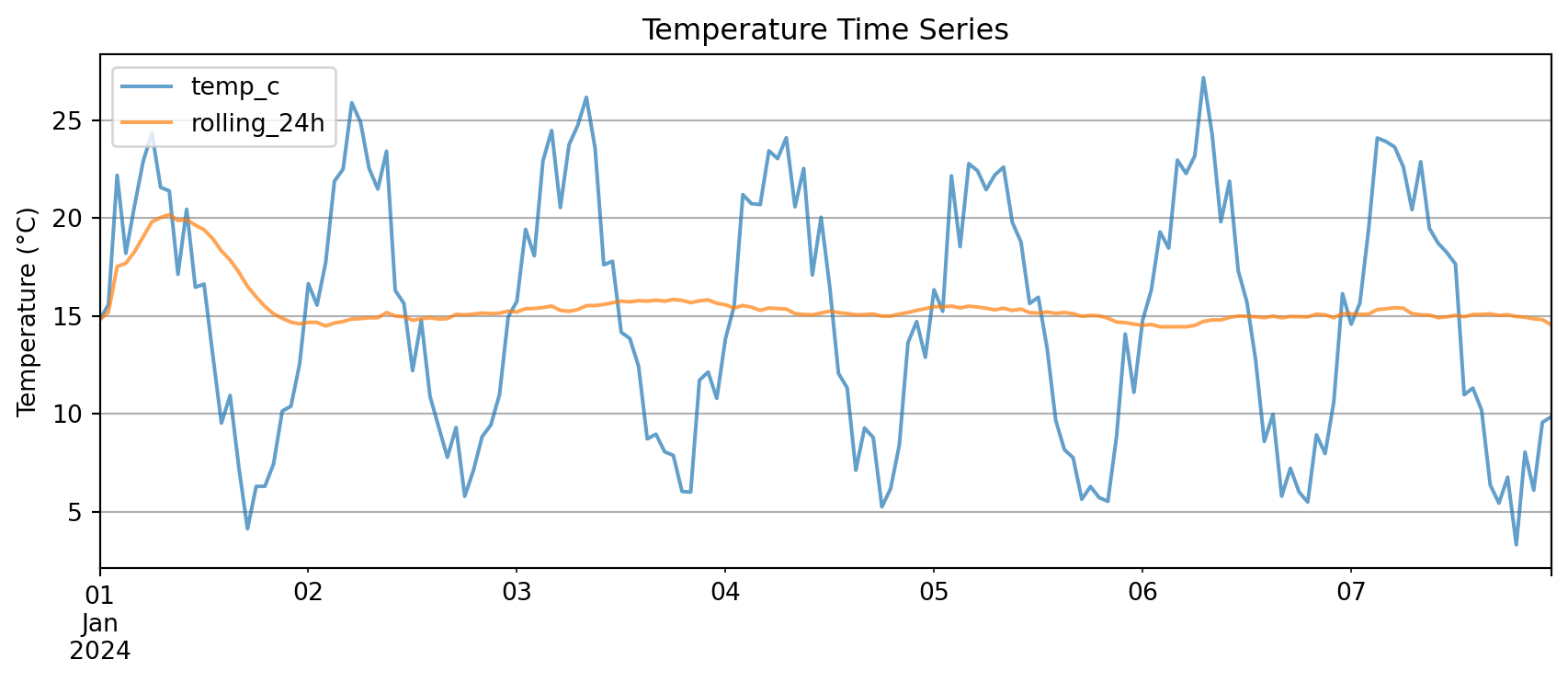
Key advantage: .plot() automatically uses the index as x-axis
Multi-Panel Time Series
For complex analysis, use matplotlib subplots:
# Create comprehensive dataset
dates = pd.date_range('2024-01-01', periods=168, freq='h')
weather = pd.DataFrame({
'temp_c': 15 + 8 * np.sin(np.arange(168) * 2 * np.pi / 24) + np.random.randn(168) * 2,
'rh_pct': 60 + 20 * np.sin(np.arange(168) * 2 * np.pi / 24 + np.pi/4) + np.random.randn(168) * 5,
'precip_mm': np.random.exponential(0.3, 168)
}, index=dates)
# Add derived quantities
weather['temp_rolling_24h'] = weather['temp_c'].rolling('24h').mean()
weather['cumulative_precip'] = weather['precip_mm'].cumsum()
# Create multi-panel plot
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, figsize=(9, 7), sharex=True)
# Panel 1: Temperature
ax1.plot(weather.index, weather['temp_c'], alpha=0.4, label='Hourly Temp')
ax1.plot(weather.index, weather['temp_rolling_24h'], linewidth=2, label='24-h Rolling Mean')
ax1.set_ylabel('Temperature (°C)')
ax1.set_title('Weather Time Series Analysis')
ax1.legend(loc='best')
ax1.grid(True, alpha=0.3)
# Panel 2: Relative Humidity
ax2.plot(weather.index, weather['rh_pct'], color='green', alpha=0.7)
ax2.set_ylabel('Relative Humidity (%)')
ax2.grid(True, alpha=0.3)
# Panel 3: Precipitation (dual y-axes)
ax3.bar(weather.index, weather['precip_mm'], width=0.04, alpha=0.6, label='Hourly Precip')
ax3_cum = ax3.twinx()
ax3_cum.plot(weather.index, weather['cumulative_precip'], color='steelblue',
linewidth=2, label='Cumulative')
ax3.set_xlabel('Date')
ax3.set_ylabel('Hourly Precip (mm)')
ax3_cum.set_ylabel('Cumulative (mm)')
ax3.legend(loc='upper left')
ax3_cum.legend(loc='upper right')
ax3.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()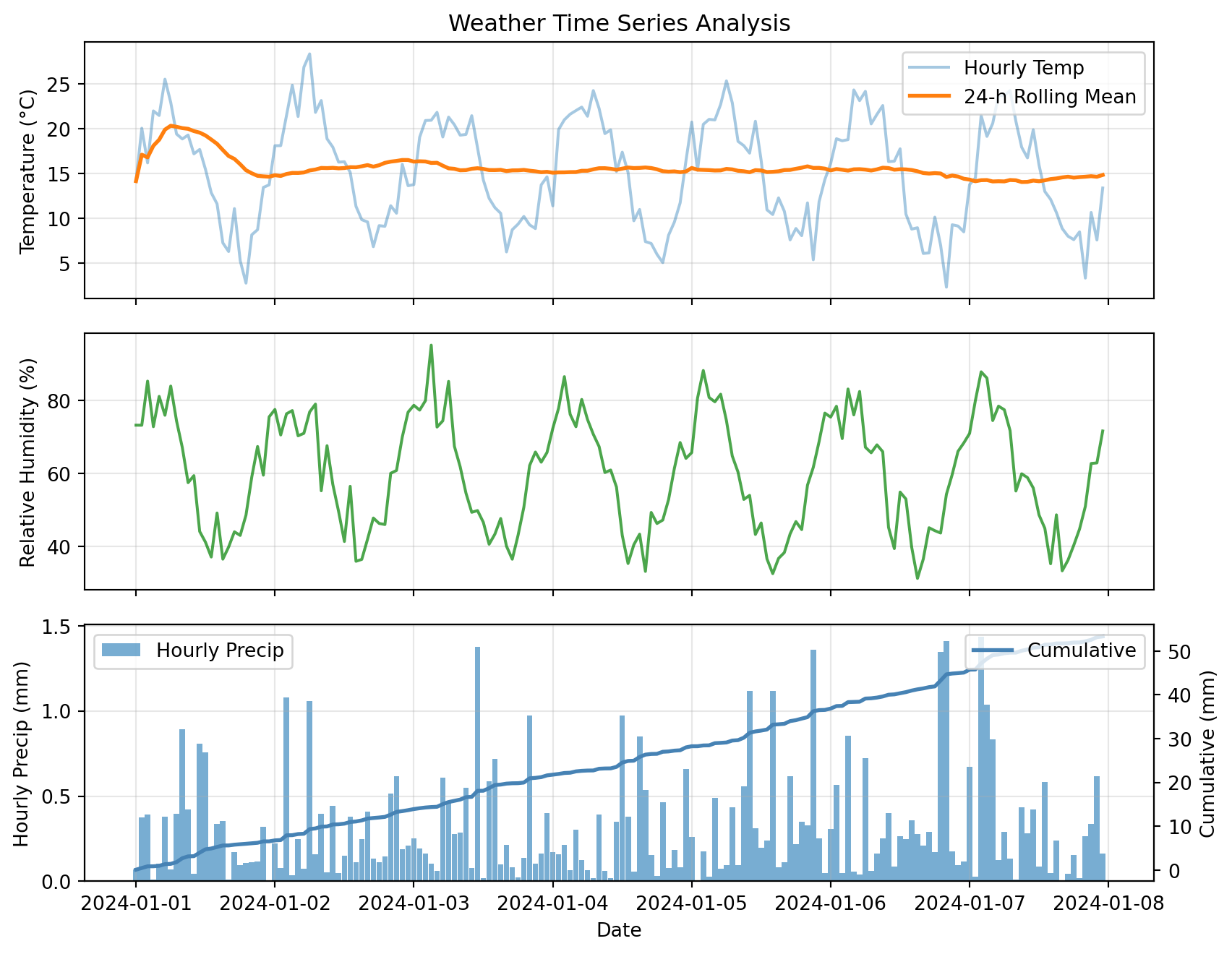
This plot combines:
- Raw and smoothed time series (top)
- Secondary variables (middle)
- Dual y-axes for different scales (bottom)
Advanced Topics
Boolean Filtering with Time Index
With a time index, boolean masks work just like NumPy:
# Create precipitation data
dates = pd.date_range('2024-01-01', periods=168, freq='h')
precip_df = pd.DataFrame({
'precip_mm': np.random.exponential(0.3, 168)
}, index=dates)
# Boolean mask: rainy hours (> 0.5 mm)
rainy_hours = precip_df['precip_mm'] > 0.5
# Count and extract
print(f"Number of rainy hours: {rainy_hours.sum()}")
print(f"\nRainiest hours:")
print(precip_df[rainy_hours].nlargest(5, 'precip_mm'))Number of rainy hours: 25
Rainiest hours:
precip_mm
2024-01-04 19:00:00 1.327211
2024-01-06 00:00:00 1.252251
2024-01-02 07:00:00 1.108433
2024-01-07 13:00:00 1.101335
2024-01-03 07:00:00 1.081913Helper Functions for Reusable Analysis
Instead of copy-pasting analysis code, wrap it in functions:
def summarize_period(df, freq='1D', temp_col='temp_c', precip_col='precip_mm'):
"""
Resample time series to specified frequency.
Parameters
----------
df : pd.DataFrame
Input dataframe with time index
freq : str
Resample frequency ('1h', '1D', '1W', etc.)
temp_col : str
Name of temperature column
precip_col : str
Name of precipitation column
Returns
-------
pd.DataFrame
Resampled data with mean temp and total precip
"""
# Check that required columns exist
if temp_col not in df.columns:
raise ValueError(f"Column '{temp_col}' not found in DataFrame")
if precip_col not in df.columns:
raise ValueError(f"Column '{precip_col}' not found in DataFrame")
# Resample with appropriate aggregations
summary = df.resample(freq).agg({
temp_col: ['mean', 'min', 'max'],
precip_col: 'sum'
})
return summary
# Test the function
daily_summary = summarize_period(weather, freq='1D', temp_col='temp_c', precip_col='precip_mm')
print(daily_summary) temp_c precip_mm
mean min max sum
2024-01-01 15.512757 5.751047 23.750560 8.726125
2024-01-02 15.340271 6.867857 26.226485 7.368808
2024-01-03 14.639162 6.676132 24.163893 7.992706
2024-01-04 15.446802 5.069988 23.905186 5.919963
2024-01-05 15.317188 3.922328 24.961065 7.447464
2024-01-06 15.226282 3.819533 24.132375 7.843172
2024-01-07 14.655429 4.862615 24.407213 8.087786Benefits:
- Reusable across projects
- Easy to test and debug
- One place to update logic
- Handles errors gracefully
Common Error: Missing Values in Rolling
Predict the output:
2024-01-01 00:00:00 NaN
2024-01-01 01:00:00 NaN
2024-01-01 02:00:00 NaN ← Missing value propagates!
2024-01-01 03:00:00 NaN
2024-01-01 04:00:00 20.5By default, NaN in window → NaN result
The Fix:
When to Use Each Tool
Decision guide:
| Goal | Tool | Example |
|---|---|---|
| Change frequency | .resample() |
Hourly → daily |
| Smooth noisy data | .rolling().mean() |
Remove high-freq noise |
| Total accumulated | .cumsum() |
Total rainfall since Jan 1 |
| Deviation from normal | Anomaly (subtract baseline) | Temp - climatology |
| Find extreme periods | Boolean mask + filter | Hours where temp > 35°C |
| Compare different aggregations | .resample().agg({...}) |
Daily mean temp, total precip |
Summary & Resources
Key Concepts Review
1. Pandas gives us labeled tables for mixed data types
- Series (1D) and DataFrames (2D)
- Named columns and time indexes
2. Reading CSVs: use parse_dates and index_col
- Converts strings to datetime objects
- Sets time index for time-aware operations
3. Resampling: change frequency
- Downsampling: high → low frequency
- Choose aggregation method: mean, sum, max, etc.
4. Rolling windows: smooth data
- Moving averages over time
- Same number of points (except NaNs at edges)
5. Anomalies: deviations from baseline
- Subtract climatology or daily mean
- Identifies unusual events
6. Cumulative sums: running totals
- Total accumulated precipitation, energy, etc.
Lab 06
Here is the sundowner data for the ATOC weather station.
https://sundowner.colorado.edu/weather/atoc1/
Turn in a cool plot and code via a github link, you will be graded on how cool it is. Turn in the code also (that uses pandas.)
Hint:
import pandas as pd
url = "https://sundowner.colorado.edu/weather/atoc1/wxobs20051004.txt"
df = pd.read_fwf(url, header=[0, 1], skiprows=[2])
date_col = [c for c in df.columns if c[1] == "Date"][0]
time_col = [c for c in df.columns if c[1] == "Time"][0]
t = (
df[time_col]
.astype(str)
.str.strip()
.str.replace(r"a$", "AM", regex=True)
.str.replace(r"p$", "PM", regex=True)
)
dt = pd.to_datetime(
df[date_col].astype(str).str.strip() + " " + t,
format="%m/%d/%y %I:%M%p",
errors="coerce",
)
df = df.set_index(dt).drop(columns=[date_col, time_col])
df.index.name = "datetime"
df.columns = [
"_".join([str(a).strip(), str(b).strip()]).replace(" ", "_").strip("_")
for a, b in df.columns
]
df
print('done')doneQuestions?
Contact
Prof. Will Chapman
wchapman@colorado.edu
willychap.github.io
ATOC Building, CU Boulder
Office Hours: Tu/Th 11:15-12:15p
See you next week!

ATOC 4815/5815 - Week 7